Molecular Docking
This page is about tools and techniques needed to do evaluate the binding strength of ligands and protiens.
Installing the software
Step one is getting software installed. I'm using the following software:
fPocket - Used detect cavities in a molecular structure
PyMOL - Used to visualize the structure of the molecules
AutoDoc Vina - This is the software that acutally calculates the docking strength.
Open Babel - Tools used convert between differnt file formats
fPocket - Used detect cavities in a molecular structure
PyMOL - Used to visualize the structure of the molecules
AutoDoc Vina - This is the software that acutally calculates the docking strength.
Open Babel - Tools used convert between differnt file formats
Hello Tools 1
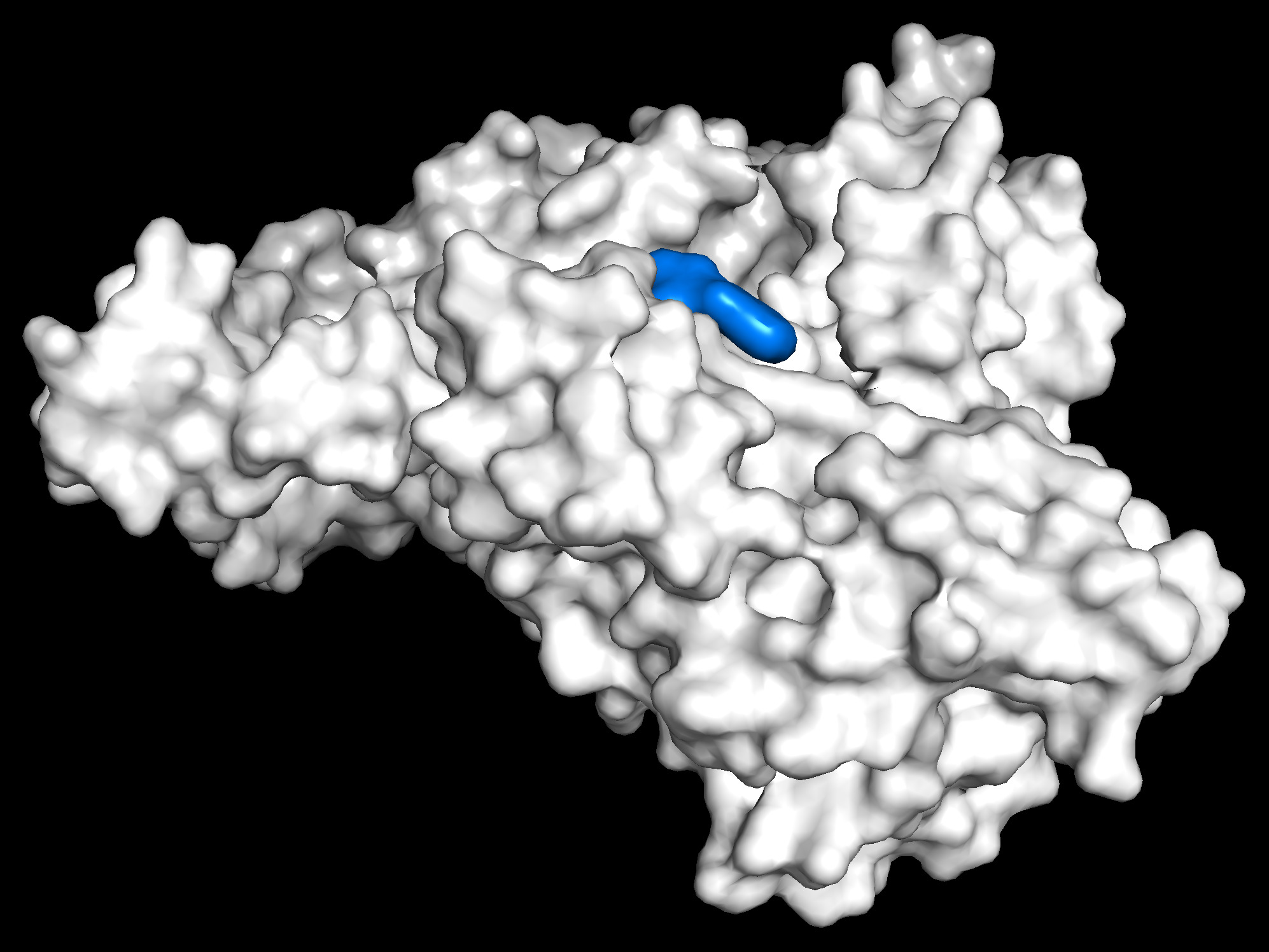
Step two is making sure the software works at the most basic level. Here we will us PyMol to view Human Serum Albumin and Wafarin before and after docking. We use Open Babel to convert the file format to a format AutoDockVina-GPU can use. Then we use AutoDockVina-GPU to perform the docking calculations.
Hello Tools 2
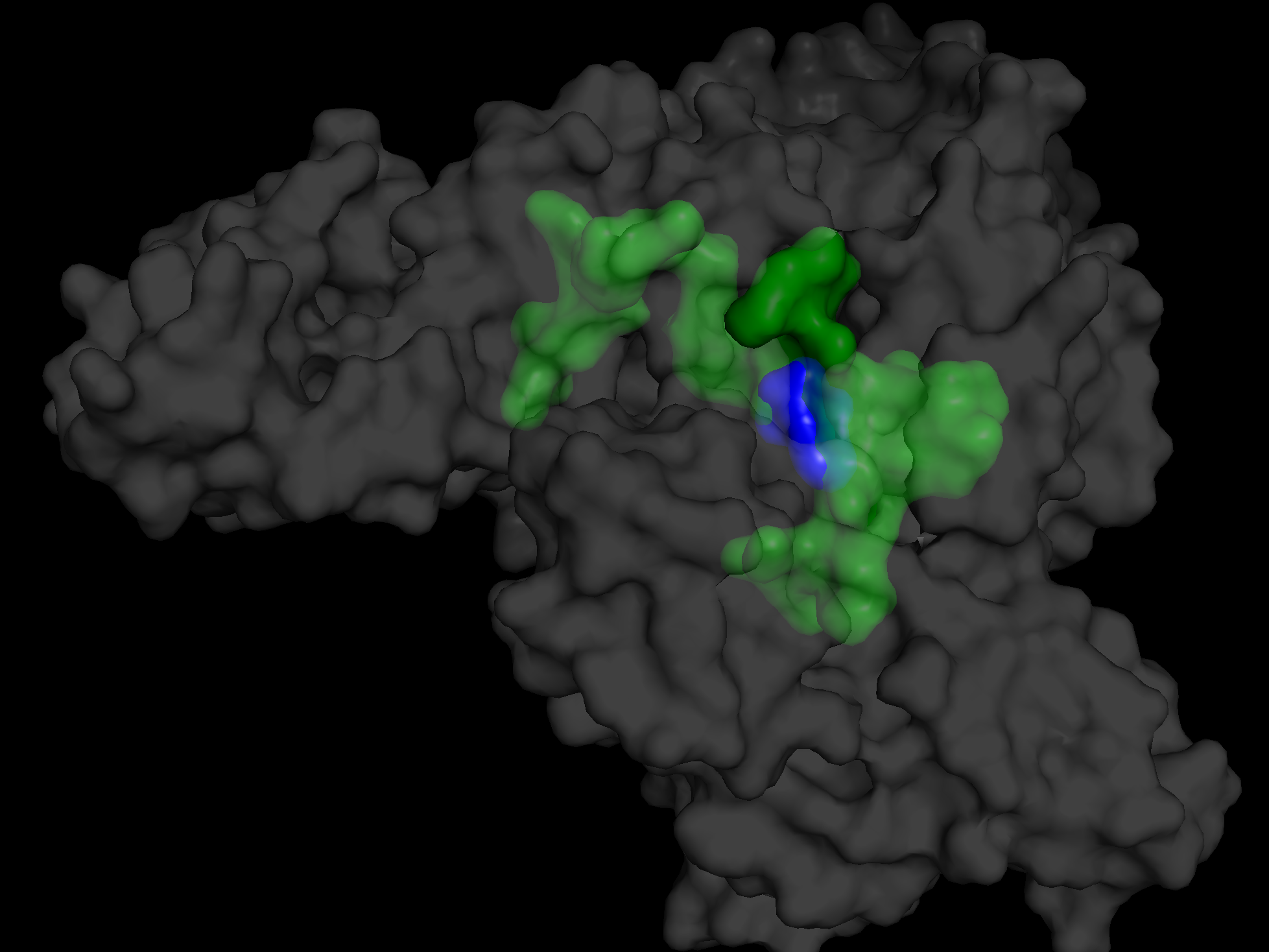
Continuing with step 2, demonstrate fPocket, basic work flow, and explination of a few of the stats generated by the tools.